|
Peak heights may vary 3-fold, which is normal. You should see evenly-spaced peaks, each with only one color. How clear are the nucleotide peaks, in general?.STEP I - Get a General Sence of How Clean the Sequence Is This document explains how to examine the normal DNA sequencing chromatogram, describing common issues and how to interpret them. Other errors can show up in the middle, invalidating individual base calls or entire swaths of data. Predictable errors occur near the beginning and again at the end of any sequencing run. That computer program, however, does make mistakes and it is the client’s responsibility to manually double-check the interpretation of the primary data. Interpretation of Sequencing ChromatogramsĪutomated DNA Sequencers generate a four-color chromatogram showing the results of the sequencing run, as well as a computer program's best guess at interpreting that data - a text file of sequence data. If you are using just the text data, you could be publishing data that is completely invalid! This page explains how to interpret a DNA sequencing chromatogram. (1990) Basic local alignment search tool, J Mol Biol 215, 403–410.In order to obtain good sequencing results, you MUST examine your sequencing chromatogram. (1981) Identification of common molecular subsequences, J Mol Biol 147, 195–197.Īltschul, S. (2008) Decoding of superimposed traces produced by direct sequencing of heterozygous indels, PLoS Comput Biol 4, e1000113. (2002) ShiftDetector: detection of shift mutations, Bioinformatics 18, 1137–1138.ĭmitriev, D. (1996) The Staden sequence analysis package, Mol Biotechnol 5, 233–241. 
(1997) PolyPhred: automating the detection and genotyping of single nucleotide substitutions using fluorescence-based resequencing, Nucleic Acids Res 25, 2745–2751. (2006) Automating resequencing-based detection of insertion-deletion polymorphisms, Nat Genet 38, 1457–1462. (2009) PineSAP – sequence alignment and SNP identification pipeline, Bioinformatics 25, 2609–2610.īhangale, T. (2008) VarDetect: a nucleotide sequence variation exploratory tool, BMC Bioinformatics 9 Suppl 12, S9. Ngamphiw, C., Kulawonganunchai, S., Assawamakin, A., Jenwitheesuk, E., Tongsima, S. (2007) PolyScan: an automatic indel and SNP detection approach to the analysis of human resequencing data, Genome Res 17, 659–666. (2007) AutoCSA, an algorithm for high throughput DNA sequence variant detection in cancer genomes, Bioinformatics 23, 1689–1691.Ĭhen, K., McLellan, M. W., Stephens, P., Raine, K., Yates, A., Mattocks, C., Tarpey, P., Butler, A., Menzies, A., Richardson, D., Jenkinson, A., Davies, H., Edkins, S., Forbes, S., Gray, K., Greenman, C., Shepherd, R., Stratton, M. Error probabilities, Genome Res 8, 186–194.ĭicks, E., Teague, J. (1998) Base-calling of automated sequencer traces using Phred. (2005) SeqDoC: rapid SNP and mutation detection by direct comparison of DNA sequence chromatograms, BMC Bioinformatics 6, 133.Įwing, B., Green, P. (2005) InSNP: a tool for automated detection and visualization of SNPs and InDels, Hum Mutat 26, 11–19.Ĭrowe, M. Manaster, C., Zheng, W., Teuber, M., Wachter, S., Doring, F., Schreiber, S., Hampe, J. (2005) novoSNP, a novel computational tool for sequence variation discovery, Genome Res 15, 436–442. Weckx, S., Del-Favero, J., Rademakers, R., Claes, L., Cruts, M., De Jonghe, P., Van Broeckhoven, C., De Rijk, P. 
(2005) SNPdetector: a software tool for sensitive and accurate SNP detection, PLoS Comput Biol 1, e53. 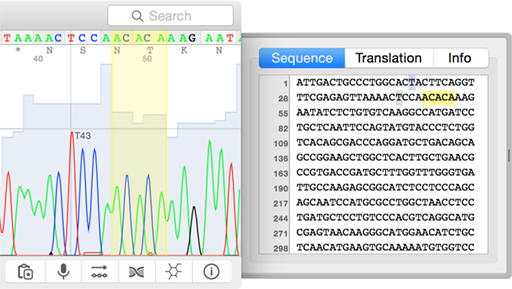
A., Yakub, I., Wei, S., Sood, R., Rowe, W., Liu, P. (2003) Automated identification of single nucleotide polymorphisms from sequencing data, J Bioinform Comput Biol 1, 253–265. Takahashi, M., Matsuda, F., Margetic, N., Lathrop, M. (1999) A general approach to single-nucleotide polymorphism discovery, Nat Genet 23, 452–456. (2001) Automated fluorescent DNA sequencing on the ABI PRISM 377, Methods Mol Biol 167, 119–152. (1992) DNA sequencing with chain-terminating inhibitors.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
